R2 Day 2
More loops, simulating data, and plots
Austin Cutler
FSU
Agenda
- In today’s class, we’re going to go over the following:
- More practice with for loops
- Bootstrap practice
- Practice simulating data
- Confidence intervals with simulations
More Loops
- Let’s start with practicing nested for loops
- Individually, create a nested loop structure that prints the sixteen weeks of the semester (i.e., Week 1, Week 2, etc.) in an outer loop and then also prints the five work days (i.e., Monday, Tuesday, etc.) in an inner loop.
- If you finish this before the others, add in a third loop that prints out the semester as well (i.e., fall, spring)
- Ignore summer and winter for this, since they don’t follow the standard sixteen week structure
But wait, there’s more (loops)
- Nested for loops can also be used to fill out the cells in a matrix
- This method is very useful for conducting power analysis when you’re testing multiple treatment effect sizes and sample sizes
- To do this, you need to apply what we learned in the previous lecture to fill in each element of a matrix
Matrix and Loops Practice
- Individually, create a nested loop that creates a matrix containing a multiplication table that goes up to and includes 12x12
- In pairs, create two 6x6 matrices, then using a nested loop, perform matrix addition, storing the results as a new 6x6 matrix
- Ensure that the appropriate cells for each matrix are added together
The Bootstrap
- The bootstrap is a very nice way of estimating confidence intervals for any quantity of interest
- How this approach works is we sample our data with replacement, generating a new dataset with the same number of observations, but some of them are missing, some are repeated, and some only appear once
- We then reestimate our quantity of interest, and then we repeat this process 1000-10000 times
- This is distinct from using data simulations for estimating confidence intervals
- As we’ll discuss later, when we simulate data we will assume a certain distribution for the data
- With this approach, we make no assumptions about the distribution of the underlying data generating process
The Bootstrap (in Code)
- The functions you’ll want to be most familiar with are
sample()andslice_sample() sample()will take a sample from a vector, andslice_sample()will take a sample from a data frame- In
sample()the argument to decide how large of a sample you’ll take issize, and forslice_sample(), it isn - In both cases, we’ll want to set this equal to the size of our original data, and set the argument
replaceto be TRUE
The Bootstrap (in Code)
- Here’s an example of the bootstrap for the mean:
Pull yourself up by…
- Using the
USJudgeRatings, first estimate the minimum, mean, and maximum of each variable- You can do this either using a for loop or by using the
summarize()function from tidyverse
- You can do this either using a for loop or by using the
- For each of these quantities, estimate confidence 95% confidence intervals using the bootstrap
- Get standard 95% confidence intervals for the mean and compare them to the bootstrap confidence intervals
Simulations
Simulating Data
- In doing quantitative social science, simulating data is a very important tool in your toolkit
- Monte Carlo simulations for power
- Conducting analysis on a rare phenomenon
- Estimating confidence intervals (more ways than bootstrapping, see King, Tomz, and Wittenburg (2000))
- Randomization inference and treatment assignment mechanisms
- Creating toy data examples to explore and demonstrate problems, or as a proof of concept
- Or to teach methods to undergraduates or grad students
Simulating Data
- To simulate data, we will most often use the “r” functions from the sets of built in, distribution family functions in r
rbinom(),rnorm(),rnbinom(),rmvnorm()*, etc.
- These functions all have companion functions that give you the pdf of a given distribution for whatever data you put in,
pbinom(),pnorm(),pnbinom(),pmvnorm()*, etc.- there are also “d” functions for the density, and “q” functions for the quantiles
- These two functions come from the
mvtnormpackage, while the rest are in base R
- These two functions come from the
rbinom()
- One of the most common simulations you’ll do is simulating Bernoulli Trials for random treatment assignment
- To do this, we will use
rbinom()
[1] 1 1 0 1 0 1nis the number of observations,sizeis the number of trials per observation, andprobis the probability of a successful trial- In practice, you’ll most often be changing
norprob
- In practice, you’ll most often be changing
rbinom() practice
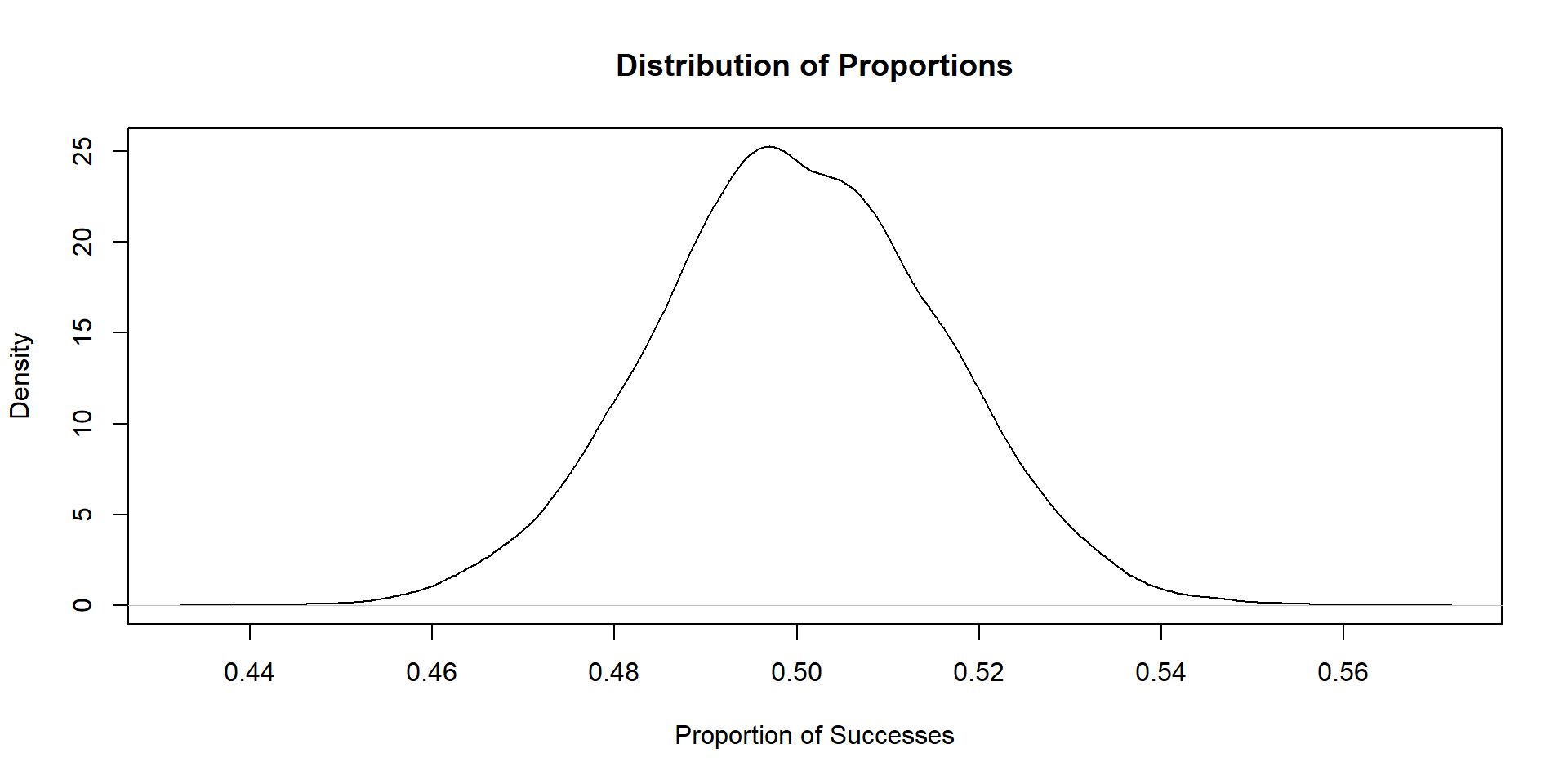
- Simulate 10,000 Bernoulli trials
- For each trial, set the number of observations to be 1000, with the probability of a “success” being .5
- For each trial, estimate the proportion of units that succeed, and store this estimate in a vector
- After all 10,000 trials, plot the distribution of that vector (using
plotorggplot)
Plot
Simulating the Normal Distribution
- We can also simulate draws from the standard normal distribution
- To do this, we use the
rnorm()function - The syntax of the code is nearly identical, but instead
rnorm()takes the argumentsmeanandsd
Practice with rnorm()
- Use a for loop to plot histograms of the standard normal distribution over 3 sample sizes: n = 10, n = 50, and n = 1000
- Repeat the previous step, but by varying the mean and standard deviation as well
- Bonus points if you incorporate the paramater value changes in your plot titles
- Reminder: plots can be stored as objects, and saved in lists
Simulating Matrices
- We can also us
rnorm()to simulate an entire matrix - An example of how we would do this is as follows:
# creating a matrix using the standard normal distribution
mat <- matrix(rnorm(4000), ncol = 4)
head(mat) [,1] [,2] [,3] [,4]
[1,] 1.6609885 0.5292288 -0.2973597 -1.0798235
[2,] 0.7807043 1.2587961 -1.3832009 -2.3535789
[3,] -0.5396074 0.6998001 0.9047782 -1.4692511
[4,] 0.2365920 -1.0059394 -0.3580972 0.9208851
[5,] 0.1890745 -0.7260650 0.3786667 -0.6581734
[6,] 0.7461223 -0.7676132 0.9537298 -0.5252005Simulating Matrices
- We can also simulate multiple distributions in a single matrix:
# creating a matrix using the standard normal distribution
mat2 <- matrix(c(rnorm(3000),rbinom(1000,1,.5)), ncol = 4)
head(mat2) [,1] [,2] [,3] [,4]
[1,] 0.48793932 0.2840526 -0.1178434 0
[2,] -0.72609351 0.8127428 0.3048668 1
[3,] 0.08315486 1.8190829 1.2636292 0
[4,] 0.21056237 0.4234619 0.3069175 0
[5,] 0.04226511 -2.2886926 1.6162669 0
[6,] 0.03433358 0.1398568 0.7513261 0Uncertainty Estimation with Monte Carlo Simulations
Multivariate Normal Distribution
- King, Tomz, and Wittenberg (2000) proposed a new method of estimating uncertainty for regression coefficients using the multivariate normal distribution
- The multivariate normal distribution is very similar to the standard normal, however \(\mu\) is now a vector of means, and instead of \(\sigma\), the variance is \(\Sigma\), which is a variance-covariance matrix
- To use this method for uncertainty estimation, you:
- Estimate your regression
- Extract the coefficients to be used as \(\mu\)
- Extract your variance-covariance matrix for \(\sigma\)
- Simulate draws from the multivariate normal
- Quantile method
Practice
- For this problem, we’re going to use the data under Day 2 on the website
- This data consists of the stats and names of each pokemon from generation 1-6, as well as how often they are used in professional tournaments (as a percentage, but numeric, i.e., 1% is 1, not 0.01)
- Your goal is to fit a linear regression model to predict tournament usage, then use the KTW method to get 95% confidence intervals.
Practice
- First, fit a linear regression model with all covariates (except for total, as you’ll run into multicolinearity issues), regressing pokemon usage onto the remaining covariates
- Extract the regression coefficients and variance covariance matrix (the
coef()andvcov()functions will help here) - Use the
rmvnorm()function from themvtnormpackage to simulate 1000 draws from the multivariate normal, using the coefficients and variance-covariance matrix from step 1 - Take the quantiles of each column of the matrix (hint: use the apply function), to get your 95% confidence intervals
Answer
Code
library(tidyverse)
pokemon_data <- read_csv('pokemon_data.csv')
fit <- lm(usage_per ~ type_1+type_2+hp+attack+defense+sp_atk+sp_def+speed+generation+legendary, data = pokemon_data)
as.data.frame(apply(mvtnorm::rmvnorm(10000, mean = coef(fit),sigma = vcov(fit)),
2,quantile, probs = c(.025,.975))) %>%
pivot_longer(cols = 1:ncol(.)) %>%
arrange(name) %>%
mutate(qi = rep(c('lower','upper'), time = nrow(.)/2)) %>%
pivot_wider(names_from = qi, values_from = value) %>%
left_join(as.data.frame(coef(fit)) %>%
mutate(name = rownames(.))) %>%
rename(beta = `coef(fit)`) %>%
select(name,beta,lower,upper) %>%
ggplot(aes(x = beta, y = name))+
geom_pointrange(aes(xmin = lower, xmax = upper))+
geom_vline(aes(xintercept = 0), linetype = 2)+
theme_classic()