[1] 4Day 3
Let’s get some lunch
Austin Cutler
FSU
Agenda
- In today’s class, we’re going to go over the following:
- The first homework assignment
- User created functions
Ifstatements- The
sandwichpackage for robust standard errors - More plotting practice
Homework check in
User created functions
The function() function
- There are some instances where in R, there isn’t a preexisting package that has a function we need
- This can happen if we have a very niche task to complete
- Fortunately, we can use
function()to make our own functions in R
function()al form
function()is incredibly flexible- In the parentheses [
()], we put in whatever arguments we’re going to give the function, and then in the brackets ({}) we put what we’re actually doing with the function - We can create conditional functions using the
iffunction andelseas well - Here is an example of a very simple user created function:
Functions Practice
- Individually, do the following:
- Simulate data from the normal distribution with 1000 observations, name this vector
x - Write a function to by hand take the mean of a variable
- Once you have the mean from your own function, use the built in mean function to check your work
Functions for Complex Calculations
- User made functions can have as many arguments as you want, and use as many other functions as you want within them
- Here is an example of a generic function:
Functions for Calculations
- We can even make user created functions for complex methods, such as SE estimation using the KTW method
- Let’s look at problem 2 from Homework 1, first we’ll load in the data and fit our model
Functions for KTW
- If you have multiple models that you’re trying to fit, it would be very beneficial to write a function to estimate KTW confidence intervals rather than doing it by hand each time
#Notice how I set some values in the function, that gives it a default of .95
ktw <- function(mod,n_sim = 1000, ci_lvl = .95, tails = 'two'){
mvtnorm::rmvnorm(n_sim,coef(mod),vcov(mod)) -> draws
if(tails == 'two'){
apply(draws, 2, quantile, probs = c((1-ci_lvl)/2,ci_lvl+(1-ci_lvl)/2)) -> ci
}
else
apply(draws, 2, quantile, probs = c(1-ci_lvl,ci_lvl)) -> ci
return(ci)
}KTW EX
- Let’s look at the summary statistics for our model
Call:
lm(formula = out_feel ~ tr, data = data)
Residuals:
Min 1Q Median 3Q Max
-13.741 -12.941 -9.741 5.907 87.059
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 12.9415 1.0363 12.488 <2e-16 ***
trpolicy 0.7991 1.4618 0.547 0.585
trpartisan 0.6059 1.4780 0.410 0.682
---
Signif. codes: 0 '***' 0.001 '**' 0.01 '*' 0.05 '.' 0.1 ' ' 1
Residual standard error: 20.54 on 1167 degrees of freedom
Multiple R-squared: 0.0002777, Adjusted R-squared: -0.001436
F-statistic: 0.1621 on 2 and 1167 DF, p-value: 0.8504- Let’s use our new function to get KTW confidence intervals
KTW EX
- We can improve this function even further, however
- While the data provided is useful, it still requires cleaning after the fact
- We can modify the original function to make improve the data we get back from the function
ktw <- function(mod,n_sim = 1000, ci_lvl = .95, tails = 'two'){
mvtnorm::rmvnorm(n_sim,coef(mod),vcov(mod)) -> draws
if(tails == 'two'){
apply(draws, 2, quantile, probs = c((1-ci_lvl)/2,ci_lvl+(1-ci_lvl)/2)) -> ci
}
else
apply(draws, 2, quantile, probs = c(1-ci_lvl,ci_lvl)) -> ci
as.data.frame(ci) %>%
pivot_longer(cols = 1:ncol(.),
names_to = 'coef') %>%
mutate(qi = rep(c('lower','upper'), each = 3)) %>%
pivot_wider(names_from = qi, values_from = value) -> ci
return(ci)
}KTW EX
# A tibble: 3 × 3
coef lower upper
<chr> <dbl> <dbl>
1 (Intercept) 10.9 15.0
2 trpolicy -1.96 3.73
3 trpartisan -2.23 3.53- Please note that this function isn’t 100% perfect
- In particular, the code for one-tailed tests only does a positive one-tailed test
- If we wanted, we could further expand this function to distinguish between positive and negative one-tailed tests
Practice Writing Functions
- Write a function called
hand_beta()that will give you the by hand regression coefficients for the model from the previous example - Create a vector that has values that range from 60 to 100, and 100 observations; name this vector a. Create a custom function called
letter_grade()that converts these numbers into a letter grade. Combine both vectors into a table. - Make a new object, x, and make it follow a normal distribution with a mean of 10 and standard deviation of 5, and 1000 observations. Create a function called
normalize()that normalizes this variable (i.e., make it have a mean of 0 and standard deviation of 1). Compare the values produced from your function to the values produced from the base R function,scale() - Make a function called
summary_stats()that returns a list of summary statistics, the mean, median, variance, and standard deviation. Apply this function to x, y, and a new variable z that follows a beta distribution (use the functionrbeta(), and set shape1 to 20 and shape2 to 5) - Write a function called
list_ext()to extract items from a list. Store the results from the previous examples in a single list, and try to extract them with your new function.
Functions to Help with graphs
- Another area where user made functions can be useful is making figures
- Hypothetically, let’s say you were working on a paper for publication and the figures required two distinct aesthetics
Custom Themes
theme_a <- function(){
theme_light()+
theme(legend.position = 'bottom',
legend.text = element_text(size = 20),
legend.title = element_text(size = 22),
panel.grid = element_line(linetype = 2,
color = alpha('lightgray',.6)),
axis.title = element_text(size = 20),
axis.text = element_text(size = 15))
}
theme_b <- function(){
theme_light()+
theme(legend.position = 'bottom',
legend.text = element_text(size = 15),
legend.title = element_text(size = 10),
panel.grid = element_line(linetype = 2,
color = alpha('lightgray',.6)),
axis.title = element_text(size = 10),
axis.text = element_text(size = 12))
}Repetitive Figures
make_coef_plot <- function(data){
data %>%
ggplot(aes(x = median, y = fct_rev(policy_id),
color = fct_rev(v_party), shape = fct_rev(v_party)))+
geom_pointrange(aes(xmin = low, xmax = high),
position = position_dodge(.5), size = .75)+
geom_vline(aes(xintercept = 0), linetype = 2)+
annotate('label', x = .486, y = 10.35, color = '#ff6666',
label = 'Party is Not Given')+
annotate('label', x = .145, y = 9.65, color = 'gray65',
label = 'Out-party')+
annotate('label', x = .08, y = 10.2, color = 'gray55',
label = 'In-party')+
#facet_wrap(vars(v_party))+
labs(x = "Effect of Race on Probability of Expecting Vignette to Adopt Own Party's Position",
y = 'Policy', color = 'Vignette Party')+
xlim(-.3,.6)+
scale_y_discrete(labels = label_wrap_gen(16))+
scale_color_manual(values = c('Not Given' = '#ff6666',
'In-party' = 'gray55',
'Out-party' = 'gray65'))+
scale_shape_manual(values = c('Not Given' = 17,
'In-party' = 19,
'Out-party' = 21))
}Sandwich
Sandwich
- The sandwich package is used to estimate robust standard errors
- Robust standard errors can be estimated for all types of models including clustered models, panel data, longitudinal data, etc.
- How this is done is by changing how we get the variance covariance matrix that is used to estimate standard errors
- More information about the package and it’s uses can be found here.
Using Sandwich
- There are several use cases for the
sandwichpackage that apply to political science work - We can use
sandwichto estimate heteroskedastic robust standard errors withvcovHC() - Or to get clustered-robust standard errors with
vcovCL() - We can even use the sandwich package to get bootstrapped standard errors with
vcovBS()- We can also add clusters to our bootstrap if needed
Using Sandwich
- The general process for using these different approaches is the same
- We want to use the function from sandwich to estimate whatever variance-covariance matrix is needed
- We then will use the
coeftest()function from thelmtestpackage to get the standard errors for based on the estimation strategy we intend on using
Practice with sandwich
- Using the model from the homework, compare the normal standard errors to heteroskedastic robust standard errors, clustered-robust standard errors, and bootstrapped standard errors
- When clustering, cluster the standard errors by 3-point partisanship (really just Republicans and Democrats since pure Independents won’t be included in the model, this variable is called
pid1) - Which method produced the largest standard error? Which one produced the smallest?
- Bonus: make a plot showing the regression coefficients with each type of confidence interval
Plot
Code
##heteroskedastic
vcov_h <- sandwich::vcovHC(fit)
fit_h <- fit_h <- lmtest::coeftest(fit, vcov = vcov_h)
##clustered-robust
vcov_cr <- sandwich::vcovCL(fit, cluster = data$pid1)
fit_cr <- lmtest::coeftest(fit, vcov = vcov_cr)
##bootstrapped
vcov_bs <- sandwich::vcovBS(fit)
fit_bs <- lmtest::coeftest(fit, vcov = vcov_bs)
##bootstrapped by hand
n_sim <- 10000
coefs <- list()
for(i in 1:n_sim){
bs_dat <- slice_sample(data, n = nrow(data), replace = TRUE)
bs_fit <- lm(out_feel ~ tr, data = bs_dat)
coefs[[i]] <- tibble(coef = names(coef(bs_fit)),
est = coef(bs_fit))
}
bind_rows(coefs) %>%
group_by(coef) %>%
reframe(est = quantile(est, probs = c(.025,.975))) %>%
mutate(qi = rep(c('lower','upper'),3)) %>%
pivot_wider(names_from = qi, values_from = est) %>%
mutate(vcov = 'Bootstrap by Hand') %>%
left_join(tibble(est = coef(fit),
coef = names(coef(fit)))) -> bs_ci
##making a figure to compare
tibble(est = coef(fit), se = fit_h[,2], vcov = 'Heteroskedastic') %>%
bind_rows(tibble(est = coef(fit), se = fit_cr[,2], vcov = 'Clustered-Robust'),
tibble(est = coef(fit), se = lmtest::coeftest(fit)[,2], vcov = 'Standard'),
tibble(est = coef(fit), se = fit_bs[,2], vcov = 'Bootstrapped')) %>%
mutate(coef = rep(row.names(fit_h), times = 4),
lower = est - 1.96*se,
upper = est + 1.96*se) %>%
bind_rows(ktw(fit) %>%
mutate(est = coef(fit),
vcov = 'KTW'),
bs_ci) %>%
mutate(vcov = factor(vcov,
levels = c('Standard',
'Heteroskedastic',
'Clustered-Robust',
'Bootstrapped',
'Bootstrap by Hand',
'KTW')),
coef = str_to_title(str_remove(coef,'tr'))) %>%
ggplot(aes(x = est, y = fct_rev(coef), color = vcov))+
geom_pointrange(aes(xmin = lower, xmax = upper),
position = position_dodge2(.45,
reverse = TRUE))+
labs(color = 'Standard Error',
y = 'Coefficient', x = 'Estimate') +
scale_color_brewer(palette = 'Set2')+
theme_b()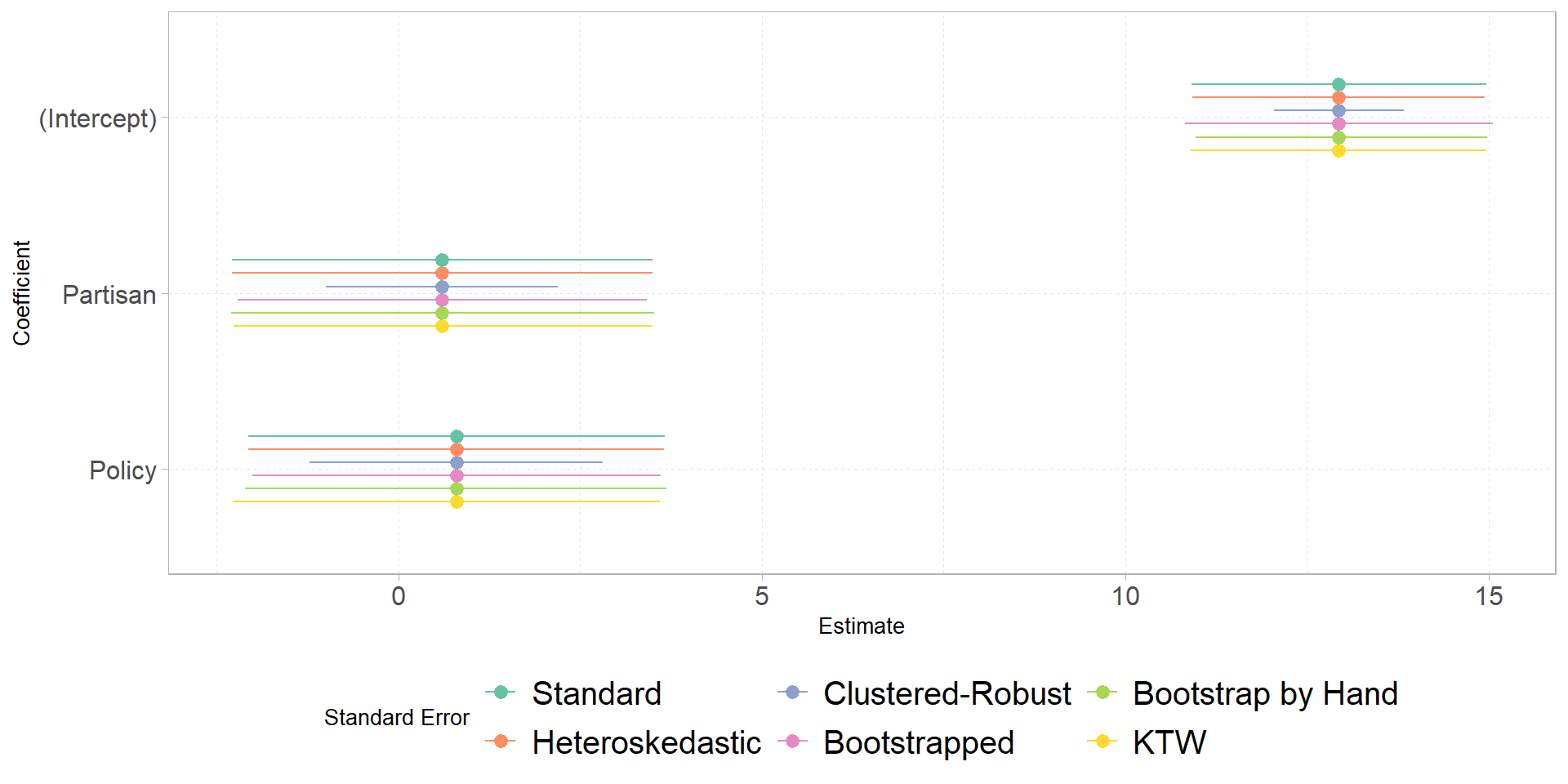
Comments
quartoto produce documents like this, you’ll want to make sure you avoid unnecessary printouts, such as warnings or messages from loaded packages, or just results that do not answer a question (e.g., printing the name of every variable for the dataset used in question 2 when only two of those variables are actually used for the assignment.)#be used for headers, you need a space between the symbol and your titleplot()when you call it